Download and run RADDOSE-3D - recommended:
- Click here for the latest RADDOSE-3D release
- Download the latest jar file
- Create a plain-text input file in the same directory as your downloaded jar file. See here for tips on writing the file.
- To run, navigate to this directory in the command prompt and use the command:
java -jar raddose3d.jar -i MyInput.txt - Help on how to run and create an input file can be found here
Additional ways to run RADDOSE-3D:
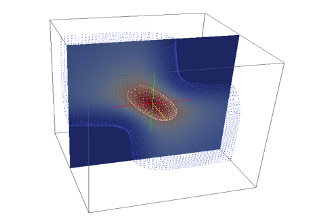
A simpler way to run RADDOSE-3D for new users, and for the automatic generation of 3D dose maps, is via the manual interface. Note that functions requiring file uploads cannot be used with the manual interface.- Click here for an online manual interface
Keep up to date with our newest source code:
RADDOSE-3D is continously in development and the Garman Group is still adding new features to our code. Our most up to date code base is openly available on GitHub. Feel free to download our newest code, run it, or even modify it.Current developments are ongoing by our team in the following areas for release in Spring 2024:
- Testing of code for estimating doses absorbed during electron diffraction experiments.
- Inclusion of the facility to enter a diffraction intensity decay model which modifies the ‘Diffraction Weighted Dose (DWD)’ output from the current ‘Fluence-Weighted Dose’ to a ‘Diffraction-Decay Weighted Dose (DDWD)’.
- Writing a User-GUI.
If you have any queries regarding these functionalities, please contact us and we can provide a working jar version of our developmental code!
User guide and help:
News:
2023-10-18
Minor fixes, including: For heavy atoms, where X-ray energies above the L3 edge but below the L1 edge, are now dealt with more accurately. This is necessary for metals in small molecule crystallography. Release available on GitHub.
2021-03-13
Minor changes, including entry of material from surrounding material for SAXS, and bug fixes. Release available on GitHub.
2019-09-20
A new major version v4.0 of RADDOSE-3D, including the ability to define a pink beam, RADDOSE-XFEL to estimate time-resolved absorbed pulses, and a new publication on the new RADDOSE-3D Monte Carlo subprogram, for simulating rotating crystals during synchrotron data collection. Release available on GitHub.
2018-08-22
When defining an oil based surrounding, the user can now define the density and elemental composition instead of the concentration. Release available on GitHub.
2018-08-14
We have a new release of the RADDOSE-3D that can now calculate fluorescent escape and simulate photoelectrons produced in the crystal and surrounding material. Source code available on GitHub.
2018-01-16
We have a new release of the RADDOSE-3D source code available on GitHub.
2017-09-16
We have a new publication on our recent developments in RADDOSE-3D.
2017-01-02
RADDOSE-3D now simulates SAXS experiments. The release is now available on Github.
2016-10-27
We have added some tips for running RADDOSE-3D, which should help for cases when the default values will give misleading/null results.
2015-06-22
RADDOSE-3D version 1.2.467 fixes an issue that caused the program to crash if heavy atoms in the solvent were not specified.
2014-12-18
We have now released the RADDOSE-3D version 1.2.427 source code at GitHub. This version also includes an updated user guide.
2014-12-11
Several improvments to functionality have been incorporated into RADDOSE-3D version 1.2.415. The absorption coefficient calculation is more accurate, particularly for structures containing nucleic acids.
2014-01-09
Together with our paper Predicting the X-ray lifetime of protein crystals we released RADDOSE-3D version 1.1.1000, the first version to support Diffraction Weighted Dose (DWD).
2013-10-20
RADDOSE-3D version 1.0.994 fixes an issue that caused a single misplaced minus sign in large CSV outputs.
2013-10-19
We have just released RADDOSE-3D version 1.0.989. This version uses a more accurate algorithm to identify the TAD-95 dose quantile, especially in very low or high dose regimes.